
ECGNet – An example application of Machine Learning for Heart Rhythm Diagnostics
TL;DR
- Deep learning applied to ECG time-series data can match expert-level arrhythmia detection performance.
- A modified 1D U-Net architecture demonstrated robust segmentation across diverse PhysioNet datasets.
- Model scaling analysis revealed that smaller architectures can maintain accuracy while reducing hardware requirements.
- Addressing data imbalance, signal variability, and demographic diversity is critical for clinical reliability.
- Lightweight AI ECG models offer a practical pathway toward deployable, assistive diagnostic systems.
Electrocardiography (ECG) remains the gold standard for non-invasive cardiac assessment, providing a vital window into the heart’s electrical health. By recording electrical impulses through surface electrodes, clinicians can identify life-critical conditions such as arrhythmias, myocardial infarctions, and conduction abnormalities.
Today, the field stands at a crossroads: the surge of digital health has moved ECG data from the clinic to the cloud. While some waveform data is now accessible via the internet and wearable devices, this data can be noisy, inconsistent, and of varying quality.
This whitepaper explores the transformative potential of Deep Learning and Artificial Intelligence in this data environment. ECGs are time-series signals, so they are well suited for convolutional neural networks, which excel at pattern recognition within sequential data. Research indicates that deep learning models can match or even exceed expert-level performance in arrhythmia detection; however, the true challenge lies in achieving this accuracy when the underlying data is uncurated or poorly annotated.
Our objective through this process was to determine how robust convolutional deep learning architectures can be when trained on representative, real-world datasets that mirror the imperfections of modern medical data collection. Our goal was to solve the problem of scale and generalizability: developing as small a model as possible that does not just work in a lab but remains clinically reliable when faced with the signal interference and class imbalances typical of diverse, internet-sourced, and poor-quality cardiac records.
AI for ECG Analysis
The application of AI to ECG signals has become an increasingly active area of research. Similar trends can be observed across other medical time-series data, such as EEG signals, which capture information from brain activity. While advances in recurrent and convolutional neural network architectures have significantly contributed to this growing interest, ECG analysis presents several additional characteristics that make it particularly attractive to researchers. These reasons include but not limited to:
- ECG waveform patterns (P-wave, QRS complex, T-wave) are highly structured, allowing neural networks to detect subtle morphological changes.
- Large public datasets (e.g., MIT-BIH Arrhythmia Database, PTB Diagnostic ECG Database, PhysioNet Challenge datasets) have enabled reproducible research.
- Deep learning models can perform classification, segmentation, denoising, and anomaly detection.
At StarFish Medical, we have significant experience with the development of tools and algorithms to manage all different types of biosignals, especially ECG. Due to the prevalence of labelled heart rhythm data, we saw an opportunity to apply ML techniques towards automated cardiac event detection and classification. ML-enabled medical devices face enhanced regulatory scrutiny when they are designed to perform a diagnostic function, however medical ML intended to alert a trained clinician to a point or points in time, without revealing an intention or diagnosis fit into an assistive, time-saving role. This niche is where medical AI shines, reducing clinician information overload, and focusing expert attention where it’s required. For this experimentation, we harnessed open-source ECG data to design and train an ML model to identify potentially aberrant cardiac events.
We opted to use a 1-D UNet, which is effective because the design captures the “big picture” by compressing the data, while using skip connections to preserve fine-grained details.
ECG Waveform patterns
ECG waveform patterns represent the heart’s electrical activity as it contracts and relaxes. A normal ECG includes the P wave (atrial depolarization), the QRS complex (rapid ventricular depolarization), and the T wave (ventricular repolarization). Variations in shape, duration, or timing can indicate cardiac abnormalities. For example, ST-segment elevation may suggest myocardial infarction, while prolonged QT intervals can signal risk of arrhythmias. Atrial fibrillation appears as irregular, fibrillatory baseline waves without distinct P waves, and ventricular tachycardia shows wide, rapid QRS complexes. Accurate interpretation helps diagnose rhythm disorders, ischemia, electrolyte imbalances, and other cardiac conditions.
Dataset Description
In the development of Machine Learning models, the quality of the underlying dataset is paramount. While we recognize that a production-ready solution eventually requires first-hand, specialized data collection, the current stage of our research utilizes four world-class open-source databases from PhysioNet. By integrating these diverse signals—ranging from healthy control groups to rare supraventricular events—we ensure our prototype is benchmarked against the same gold standards used by the global medical research community.
The MIT-BIH Arrhythmia Database contains 48 half-hour, two-channel ECG recordings sampled at 360 Hz, drawn from ambulatory Holter data. It includes a wide range of arrhythmias, with each beat annotated by expert cardiologists, making it a benchmark dataset for arrhythmia detection research. The MIT-BIH Normal Sinus Rhythm Database provides long-term ECG recordings from healthy individuals with no significant arrhythmias. These clean sinus-rhythm signals serve as a control set for comparing pathological versus normal cardiac patterns.
The MIT-BIH Supraventricular Arrhythmia Database adds 78 recordings selected to emphasize supraventricular events, such as atrial arrhythmias, that are underrepresented in the core MIT-BIH Arrhythmia dataset. These recordings include detailed beat annotations to support the study of conditions originating above the ventricles.
The St. Petersburg INCART 12-lead Arrhythmia Database consists of 75 half-hour segments extracted from 12-lead Holter recordings of patients evaluated for coronary artery disease. Sampled at 257 Hz, it offers a multi-lead perspective and contains more than 175,000 annotated beats, covering diverse ventricular and supraventricular abnormalities.
Combining these diverse datasets ensures our models are resilient and generalized, rather than over-fit to a single clinical environment. This multi-source strategy provides the rigorous benchmarking necessary for our current research phase while paving the way for our transition to proprietary data collection and clinical-grade deployment.
Prototype Description
Model Architecture, Modified 1D U-Net
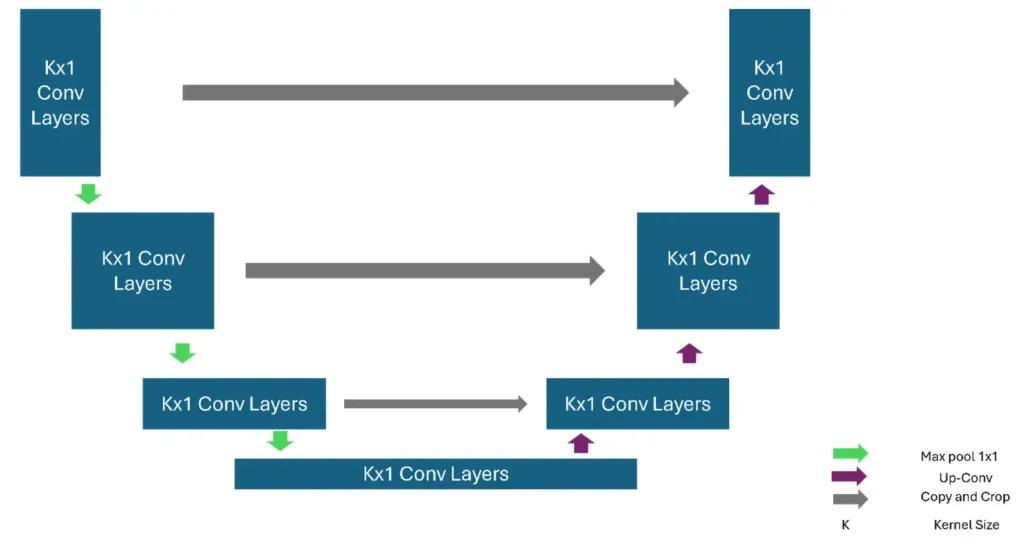
A U-Net is a convolutional neural network architecture originally proposed for biomedical image segmentation, though it has since become a foundation for many dense-prediction tasks. Its defining characteristic is the U-shaped topology composed of a contracting path (encoder) and an expanding path (decoder) connected by skip connections.
The encoder consists of repeated blocks of 3 × 3 convolutions, nonlinearities (typically ReLU), and downsampling via max pooling. As the spatial resolution decreases, the number of feature channels increases, enabling the network to learn increasingly abstract representations. The decoder mirrors this structure: each stage upsamples the feature maps, often using transposed convolutions or interpolation followed by convolution, progressively restoring spatial resolution.
A key innovation is the use of skip connections that concatenate feature maps from the encoder to corresponding decoder layers. These connections preserve high-resolution spatial information lost during downsampling and improve gradient flow during training, allowing precise localization even with relatively few training samples.
U-Net is trained end-to-end with loss functions such as cross-entropy or Dice loss, depending on class imbalance. Variants extend the architecture with attention mechanisms, residual blocks, or 3-D convolutions for volumetric data. Its efficiency, strong performance on small datasets, and ability to produce detailed segmentation maps have made U-Net a cornerstone model in medical imaging and beyond.
Class Description
As already mentioned, the model is a multiclass classifier for every beat and non-beat part of the waveform sample. Class 0 is mapped to background portions of the waveform, between heart rhythms. Class 1 refers to a section of the signal surrounding a normal beat, and Class 2 refers to the region around abnormal beats.
Using the scheme above, we can count the number of abnormal beats per patient.
Methods and Results
We trained each model for 5,000 epochs, a limit that was chosen to be a long enough to allow the many differently sized models to converge for direct comparison. Each step involved using a randomly selected two-second, single-channel chunk of ECG signal from each recording. We varied the model hyperparameters to investigate the effects of model scaling on training dynamics.
Hyperparameters
The following are the hyperparameters that we used for our 1D U-Net instances. Using these hyperparameters, we can create and compare several models and get the one that chooses the best results. Moreover, these are the most manipulated hyperparameters in convolutional neural networks.
Kernel Size is the size of the window that convolves over the time series. For image processing, the kernel is a two-dimensional set of parameters. For time-series data, which is our case, it is a one-dimensional array of parameters, and thus the kernel size is a positive integer.
The Dilation Rate is a hyperparameter that defines the spacing between kernel weights during the convolution operation. In a standard 1D convolution, the dilation rate equals 1, meaning the kernel processes adjacent samples in the time series. U-Net Depth refers to the number of successive levels in the hierarchical structure, leading from the input layer to the bottleneck.
Metrics
To comprehensively evaluate the performance of the 1D U-Net segmentation model, we selected three complementary metrics: Cross-Entropy Loss, Intersection over Union (IoU), and Macro F1-score. Cross-Entropy Loss serves as the primary training objective and measures how well the model predicts class probabilities. IoU evaluates the quality of overlap between predicted and true segments, capturing spatial or temporal alignment. Macro F1-score balances precision and recall across all classes, ensuring that minority classes are treated equally. Together, these metrics provide insight into optimization behavior, segmentation accuracy, and class-wise performance fairness. In the following paragraphs we define these metrics with more details.
Cross -entropy Loss: In the context of 1D U-Nets, Cross-Entropy Loss is the primary objective function used during training. It measures the performance of a classification model whose output is a probability value between 0 and 1.
Intersection over Union (IoU): Also known as the Jaccard Index, IoU is a spatial or temporal agreement metric that evaluates how well the predicted segment overlaps with the ground truth.
Macro F1: The F1Score is the harmonic mean of Precision and Recall. In multi-class 1D segmentation, we use the Macro version to ensure all classes are treated equally.
Model Evaluation Results
In the following table, the model names are laid out with the format M<k><d><i>, in which k is the kernel size, d is the dilation, and i is the inner depth of the model.
| Name | CE Loss | IoU | Macro F1 |
|---|---|---|---|
| M3_7_8 | 0.1120 | 0.8809 | 0.9345 |
| M3_7_32 | 0.0862 | 0.9009 | 0.9462 |
| M3_14_8 | 0.1108 | 0.8823 | 0.9354 |
| M3_14_32 | 0.0835 | 0.9042 | 0.9482 |
| M3_21_8 | 0.1047 | 0.8857 | 0.9374 |
| M3_21_32 | 0.0847 | 0.9060 | 0.9493 |
| M7_7_8 | 0.1714 | 0.9001 | 0.6186 |
| M7_7_32 | 0.0765 | 0.9129 | 0.9721 |
The trend shows that using smaller kernels with high dilation rates provides similar performance to larger kernels with smaller dilations. This indicates that relatively little information from the high -frequency domain is used by our model to classify regions of the signal. Using this knowledge, we could save significant memory in a hypothetical deployment by reducing both sampling rate and kernel dilation in tandem, unlocking the ability to pair the model with less expensive hardware, for instance in edge devices that a patient can take home.
After observing the general results and the trend, it is also informative to examine how the model outputs predictions for a given ECG sample. As shown in the image below, the prediction closely matches the ground truth.
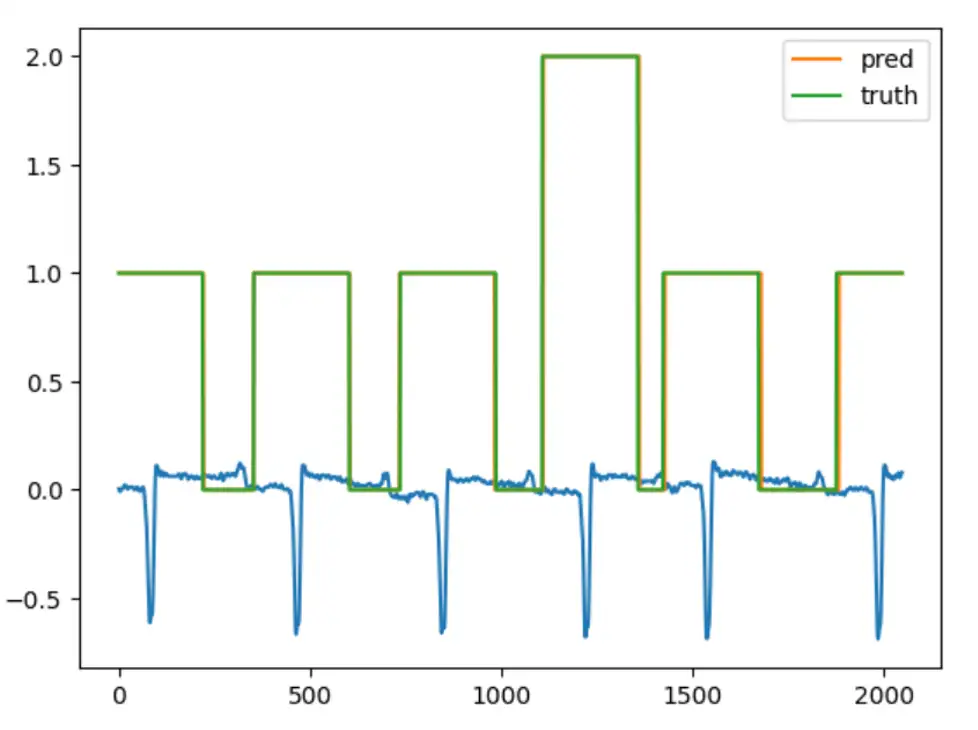
Performance Scaling Results
During our hyperparameter exploration, we identified an interesting configuration that performs far beyond what its size would suggest when compared to larger models. A model with two input features and an encoder-decoder depth of four layers achieve training dynamics like those of a much larger model.
This example highlights the benefit of expansive hyperparameter exploration; the impact of scaling is rarely linear, and the ideal balance of size and performance can exist where one least expects it.
Addressing Barriers in AI Diagnostics
Achieving clinical-grade reliability requires overcoming hurdles in data integrity and diversity. The primary challenge lies in Data Scarcity and Fragmentation; high-quality ECG datasets are frequently siloed within proprietary hospital networks, making it difficult to aggregate the volume of data needed for robust training. When data is available, it is often highly imbalanced, as healthy heart rhythms naturally outnumber rare pathologies. To combat this, we utilize strategic class reweighting and the clustering of rare morphologies to ensure the model remains sensitive to critical, less frequent events.
Furthermore, signal variability remains a technical roadblock. ECGs recorded on different devices often vary in sampling frequency and lead placement. We mitigate this through data normalization and resampling techniques to create a universal signal language. Finally, we account for demographic shifts; factors such as age, biological sex, and pre-existing conditions can subtly alter ECG morphology. By curating diverse training sets that reflect a broad patient population, we ensure our model generalizes accurately across all demographics, moving beyond average results toward personalized clinical reliability.
Roadmap for Innovation: The Future of AI-Driven ECG Analysis
As we look toward the next generation of cardiac diagnostics, our development roadmap focuses on transitioning from binary detection to a more comprehensive understanding of heart health. The application of deep learning techniques to biological time -series analysis can improve clinical efficiency and reduce cognitive load by decreasing the volume of noise a clinician must sift through to identify signals of interest. Our example cardiac U-Net demonstrates a model that is lightweight enough to function on a small, inexpensive, non-networked ECG recording system while still serving as an effective clinical aid.
The future of AI in medical devices appears particularly promising in time-saving applications where models provide improved information to support faster and clearer clinical decision-making. And Starfish Medical is well positioned to respond to emerging opportunities and help clients advance their development efforts.
References
Liu X, Wang H, Li Z, Qin L. Deep learning in ECG diagnosis: A review. Knowledge-Based Systems. 2021 Sep 5;227:107187.
Pyakillya B, Kazachenko N, Mikhailovsky N. Deep learning for ECG classification. InJournal of physics: conference series 2017 Oct 1 (Vol. 913, No. 1, p. 012004). IOP Publishing.
Murat F, Yildirim O, Talo M, Baloglu UB, Demir Y, Acharya UR. Application of deep learning techniques for heartbeats detection using ECG signals-analysis and review. Computers in biology and medicine. 2020 May 1;120:103726.
Dataset sources
MIT-BIH Arrhythmia Database v1.0.0
MIT-BIH Normal Sinus Rhythm Database v1.0.0
MIT-BIH Supraventricular Arrhythmia Database v1.0.0
St Petersburg INCART 12-lead Arrhythmia Database v1.0.0
Ronneberger, O., Fischer, P. and Brox, T., 2015, October. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical image computing and computer-assisted intervention (pp. 234-241). Cham: Springer international publishing.
// Add one D Unet
Ali Bahrani is full-stack and systems software engineer at StarFish Medical with 5+ years of experience building high-performance, safety-critical software for medical and financial systems. Specialized in C++ and cloud-native architectures, delivering production-grade applications, DevOps pipelines, and ML-powered services. Experienced in Agile and regulated environments (ISO 13485), with a strong focus on reliability, testing, and modernizing legacy systems.
Thor Tronrud, PhD, is a Machine Learning Scientist at StarFish Medical who specializes in the development and application of machine learning tools. Previously an astrophysicist working with magnetohydrodynamic simulations, Thor joined StarFish in 2021 and has applied machine learning techniques to problems including image segmentation, signal analysis, and language processing.
Title Image: Adobe Stock
Related Resources

Nick and Nigel explore the science behind hand sanitizer formulations. They discuss how alcohol interacts with bacterial cells, why water improves its effectiveness, and what the additional ingredients in sanitizer actually do.

Scott Phillips, CEO of StarFish Medical, sits down with Peter van der Velden, Managing General Partner at Lumira Ventures, to explore the strategic thinking behind major MedTech transactions and investments.

Computer vision technologies such as convolutional neural networks and vision transformers are transforming how AI analyzes medical images, each offering distinct advantages depending on the application and computing environment.

Ariana and Mark explore how prototype strategy helps teams reduce technical risk and accelerate progress.